ParaView/Users Guide/VTK Data Model
Introduction
To use ParaView effectively, you need to understand the ParaView data model. This chapter briefly introduces the VTK data model used by ParaView. For more details, refer to one of the VTK books.
The most fundamental data structure in VTK is a data object. Data objects can either be scientific datasets such rectilinear grids or finite elements meshes (see below) or more abstract data structures such as graphs or trees. These datasets are formed of smaller building blocks: mesh (topology and geometry) and attributes.
Mesh
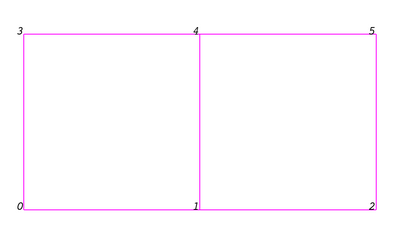
Even though the actual data structure used to store the mesh in memory depends on the type of the dataset, some abstractions are common to all types. In general, a mesh consists of vertices (points) and cells (elements, zones). Cells are used to discretize a region and can have various types such a tetrahedra, hexahedra etc. Each cell contains a set of vertices. The mapping from cells to vertices is called the connectivity. Note that even though it is possible to define data elements such as faces and edges, VTK does not represent these explicitly. Rather, they are implied by a cell's type and its connectivity. One exception to this rule is the arbitrary polyhedron which explicitly stores its faces. Figure 3.1 is an example mesh that consists of 2 cells. The first cell is defined by vertices (0, 1, 3, 4) and the second cell is defined by vertices (1, 2, 4, 5). These cells are neighbors because they share the edge defined by the points (1, 4).
A mesh is fully defined by its topology and the spatial coordinates of its vertices. In VTK, the point coordinates may be implicit or explicitly defined by a data array of dimensions (number_of_points x 3).
Attributes (fields, arrays)
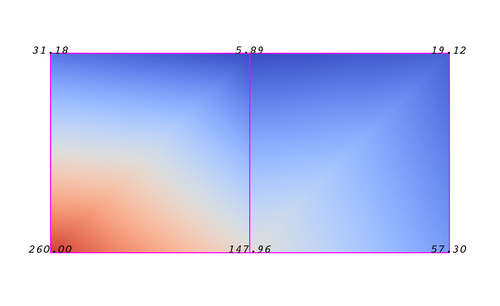
An attribute (or a data array or field) defines the discrete values of a field over the mesh. Examples of attributes include pressure, temperature, velocity and stress tensor. Note that VTK does not specifically define different types of attributes. All attributes are stored as data arrays which can have an arbitrary number of components. ParaView makes some assumptions in regards to the number of components. For example, a 3-component array is assumed to be an array of vectors. Attributes can be associated with points or cells. It is also possible to have attributes that are not associated with either. Figure 3.2 demonstrates the use of a point-centered attribute. Note that the attribute is only defined on the vertices. Interpolation is used to obtain the values everywhere else. The interpolation functions used depend on the cell type. See VTK documentation for details.
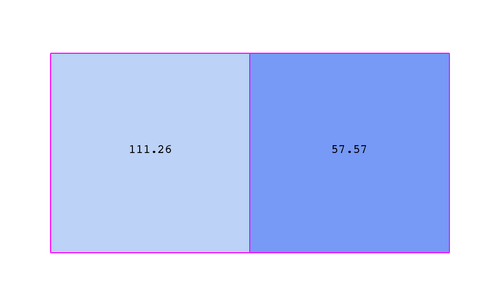
Figure 3.3 demonstrates the use of a cell-centered attribute. Note that cell-centered attributes are assumed to be constant over each cell. Due to this property, many filters in VTK cannot be directly applied to cell-centered attributes. It is normally required to apply a Cell Data to Point Data filter. In ParaView, this filter is applied automatically when necessary.
Uniform Rectilinear Grid (Image Data)
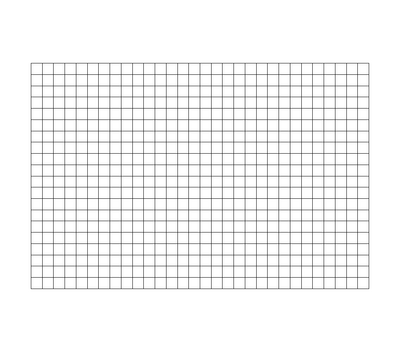
A uniform rectilinear grid, or image data, defines its topology and point coordinates implicitly. To fully define the mesh for an image data, VTK uses the following:
- Extents - these define the minimum and maximum indices in each direction. For example, an image data of extents (0, 9), (0, 19), (0, 29) has 10 points in the x-direction, 20 points in the y-direction and 30 points in the z-direction. The total number of points is 10*20*30.
- Origin - this is the position of a point defined with indices (0, 0, 0)
- Spacing - this is the distance between each point. Spacing for each direction can defined independently
The coordinate of each point is defined as follows: coordinate = origin + index*spacing where coordinate, origin, index and spacing are vectors of length 3.
Note that the generic VTK interface for all datasets uses a flat index. The (i,j,k) index can be converted to this flat index as follows: idx_flat = k*(npts_x*npts_y) + j*nptr_x + i.
A uniform rectilinear grid consists of cells of the same type. This type is determined by the dimensionality of the dataset (based on the extents) and can either be vertex (0D), line (1D), pixel (2D) or voxel (3D).
Due to its regular nature, image data requires less storage than other datasets. Furthermore, many algorithms in VTK have been optimized to take advantage of this property and are more efficient for image data.
Rectilinear Grid
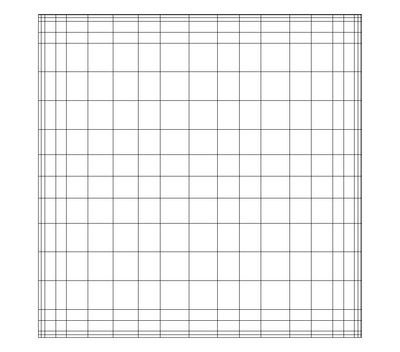
A rectilinear grid such as Figure 3.5 defines its topology implicitly and point coordinates semi-implicitly. To fully define the mesh for a rectilinear grid, VTK uses the following:
- Extents - these define the minimum and maximum indices in each direction. For example, a rectilinear grid of extents (0, 9), (0, 19), (0, 29) has 10 points in the x-direction, 20 points in the y-direction and 30 points in the z-direction. The total number of points is 10*20*30.
- Three arrays defining coordinates in the x-, y- and z-directions. These arrays are of length npts_x, npts_y and npts_z. This is a significant savings in memory as total memory used by these arrays is npts_x+npts_y+npts_z rather than npts_x*npts_y*npts_z.
The coordinate of each point is defined as follows: coordinate = (coordinate_array_x(i), coordinate_array_y(j), coordinate_array_z(k))".
Note that the generic VTK interface for all datasets uses a flat index. The (i,j,k) index can be converted to this flat index as follows: idx_flat = k*(npts_x*npts_y) + j*nptr_x + i.
A rectilinear grid consists of cells of the same type. This type is determined by the dimensionality of the dataset (based on the extents) and can either be vertex (0D), line (1D), pixel (2D) or voxel (3D).
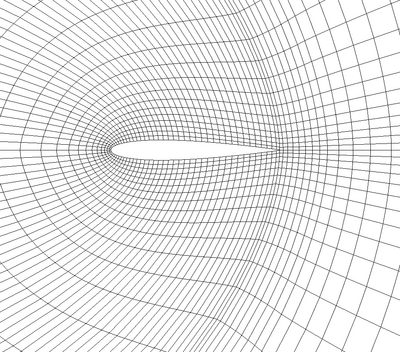
Curvilinear Grid (Structured Grid)
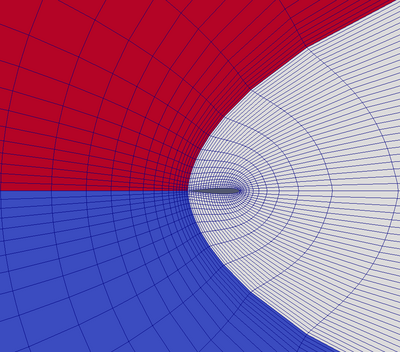
A curvilinear grid, such as Figure 3.6, defines its topology implicitly and point coordinates explicitly. To fully define the mesh for a curvilinear grid, VTK uses the following:
- Extents - these define the minimum and maximum indices in each direction. For example, a curvilinear grid of extents (0, 9), (0, 19), (0, 29) has 10*20*30 points regularly defined over a curvilinear mesh.
- An array of point coordinates. This arrays stores the position of each vertex explicitly.
The coordinate of each point is defined as follows: coordinate = coordinate_array(idx_flat)". The (i,j,k) index can be converted to this flat index as follows: idx_flat = k*(npts_x*npts_y) + j*nptr_x + i.
A curvilinear grid consists of cells of the same type. This type is determined by the dimensionality of the dataset (based on the extents) and can either be vertex (0D), line (1D), quad (2D) or hexahedron (3D).
AMR Dataset
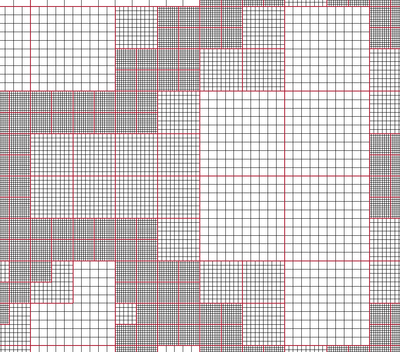
VTK natively support Berger-Oliger type AMR (Adaptive Mesh Refinement) datasets, as shown in Figure 3.7. An AMR dataset is essentially a collection of uniform rectilinear grids grouped under increasing refinement ratios (decreasing spacing). VTK's AMR dataset does not force any constraint on whether and how these grids should overlap. However, it provides support for masking (blanking) sub-regions of the rectilinear grids using an array of bytes. This allows VTK to process overlapping grids with minimal artifacts. VTK can automatically generate the masking arrays for Berger-Oliger compliant meshes.
Unstructured Grid
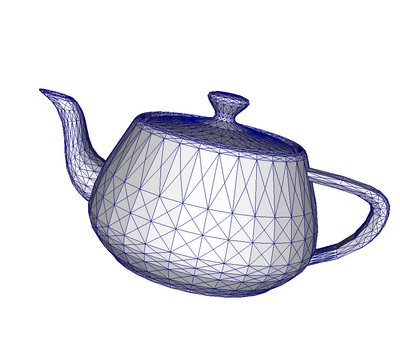
An unstructured grid such as Figure 3.8 is the most general primitive dataset type. It stores topology and point coordinates explicitly. Even though VTK uses a memory-efficient data structure to store the topology, an unstructured grid uses significantly more memory to represent its mesh. Therefore, use an unstructured grid only when you cannot represent your dataset as one of the above datasets. VTK supports a large number of cell types, all of which can exist (heterogeneously) within one unstructured grid. The full list of all cell types supported by VTK can be found in the file vtkCellType.h in the VTK source code. Here is the list as of when this document was written:
| VTK_EMPTY_CELL | VTK_VERTEX |
| VTK_POLY_VERTEX | VTK_LINE |
| VTK_POLY_LINE | VTK_TRIANGLE |
| VTK_TRIANGLE_STRIP | VTK_POLYGON |
| VTK_PIXEL | VTK_QUAD |
| VTK_TETRA | VTK_VOXEL |
| VTK_HEXAHEDRON | VTK_WEDGE |
| VTK_PYRAMID | VTK_PENTAGONAL_PRISM |
| VTK_HEXAGONAL_PRISM | VTK_QUADRATIC_EDGE |
| VTK_QUADRATIC_TRIANGLE | VTK_QUADRATIC_QUAD |
| VTK_QUADRATIC_TETRA | VTK_QUADRATIC_HEXAHEDRON |
| VTK_QUADRATIC_WEDGE | VTK_QUADRATIC_PYRAMID |
| VTK_BIQUADRATIC_QUAD | VTK_TRIQUADRATIC_HEXAHEDRON |
| VTK_QUADRATIC_LINEAR_QUAD | VTK_QUADRATIC_LINEAR_WEDGE |
| VTK_BIQUADRATIC_QUADRATIC_WEDGE | VTK_BIQUADRATIC_QUADRATIC_HEXAHEDRON |
| VTK_BIQUADRATIC_TRIANGLE | VTK_CUBIC_LINE |
| VTK_CONVEX_POINT_SET | VTK_POLYHEDRON |
| VTK_PARAMETRIC_CURVE | VTK_PARAMETRIC_SURFACE |
| VTK_PARAMETRIC_TRI_SURFACE | VTK_PARAMETRIC_QUAD_SURFACE |
| VTK_PARAMETRIC_TETRA_REGION | VTK_PARAMETRIC_HEX_REGION |
Many of these cell types are straightforward. For details, see VTK documentation.
Polygonal Grid (Polydata)
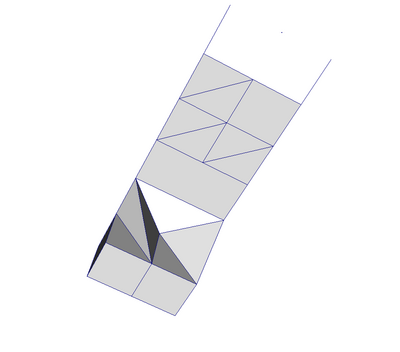
A polydata such as Figure 3.9 is a specialized version of an unstructured grid designed for efficient rendering. It consists of 0D cells (vertices and polyvertices), 1D cells (lines and polylines) and 2D cells (polygons and triangle strips). Certain filters that generate only these cell types will generate a polydata. Examples include the Contour and Slice filters. An unstructured grid, as long as it has only 2D cells supported by polydata, can be converted to a polydata using the Extract Surface filter. A polydata can be converted to an unstructured grid using Clean to Grid.
Table
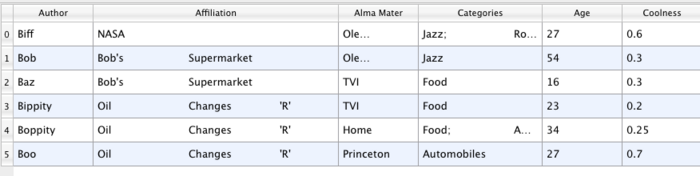
A table like Table 3.1 is a tabular dataset that consists of rows and columns. All chart views have been designed to work with tables. Therefore, all filters that can be shown within the chart views generate tables. Also, tables can be directly loaded using various file formats such as the comma separated values format. Tables can be converted to other datasets as long as they are of the right format. Filters that convert tables include Table to Points and Table to Structured Grid.
Multiblock Dataset
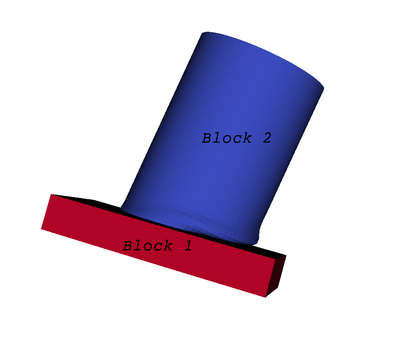
You can think of a multi-block dataset as a tree of datasets where the leaf nodes are "simple" datasets. All of the data types described above, except AMR, are "simple" datasets. Multi-block datasets are used to group together datasets that are related. The relation between these datasets is not necessarily defined by ParaView. A multi-block dataset can represent an assembly of parts or a collection of meshes of different types from a coupled simulation. Multi-block datasets can be loaded or created within ParaView using the Group filter. Note that the leaf nodes of a multi-block dataset do not all have to have the same attributes. If you apply a filter that requires an attribute, it will be applied only to blocks that have that attribute.
Multipiece Dataset
Multi-piece datasets such as Figure 3.11 are similar to multi-block datasets in that they group together simple datasets with one key difference. Multi-piece datasets group together datasets that are part of a whole mesh - datasets of the same type and with same attributes. This data structure is used collect datasets produced by a parallel simulation without having to append the meshes together. Note that there is no way to create a multi-piece dataset within ParaView, but only by using certain readers. Furthermore, multi-piece datasets act, for the most part, as simple datasets. For example, it is not possible to extract individual pieces or obtain information about them.